Northen Lab researchers Ying Wang, Tami Swenson, Anita Silver, Peter Andeer, Amber Golini, Suzanne Kosina, Benjamin Bowen, and Trent Northen are authors on the new paper “Substrate Utilization and Competitive Interactions Among Soil Bacteria Vary With Life-History Strategies.” This paper looks at genomic traits, exometabolite profiles, and interactions among soil bacteria that are copiotrophic (adapted to resource-rich environments) and oligotrophic (adapted to resource-poor conditions).
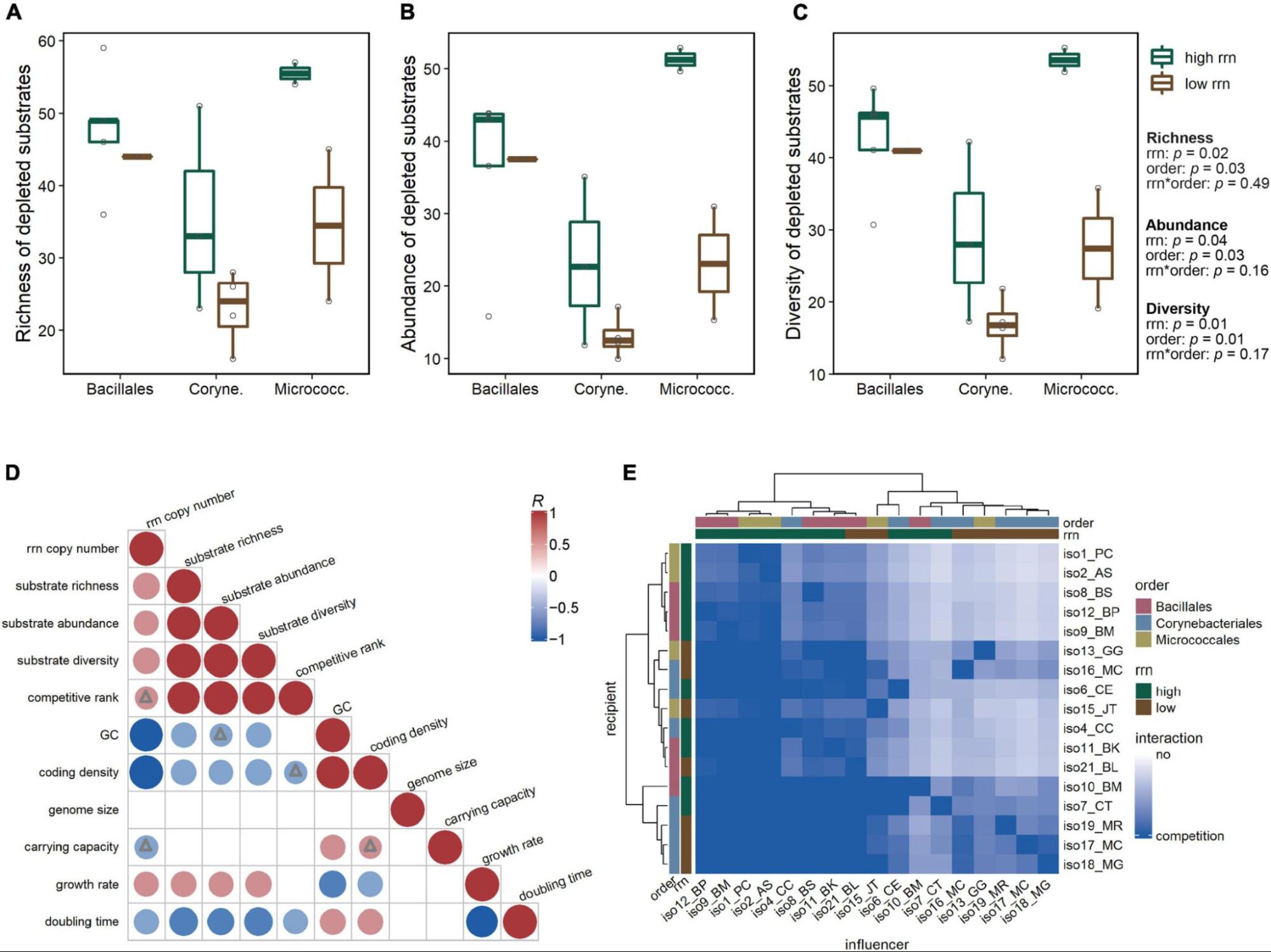
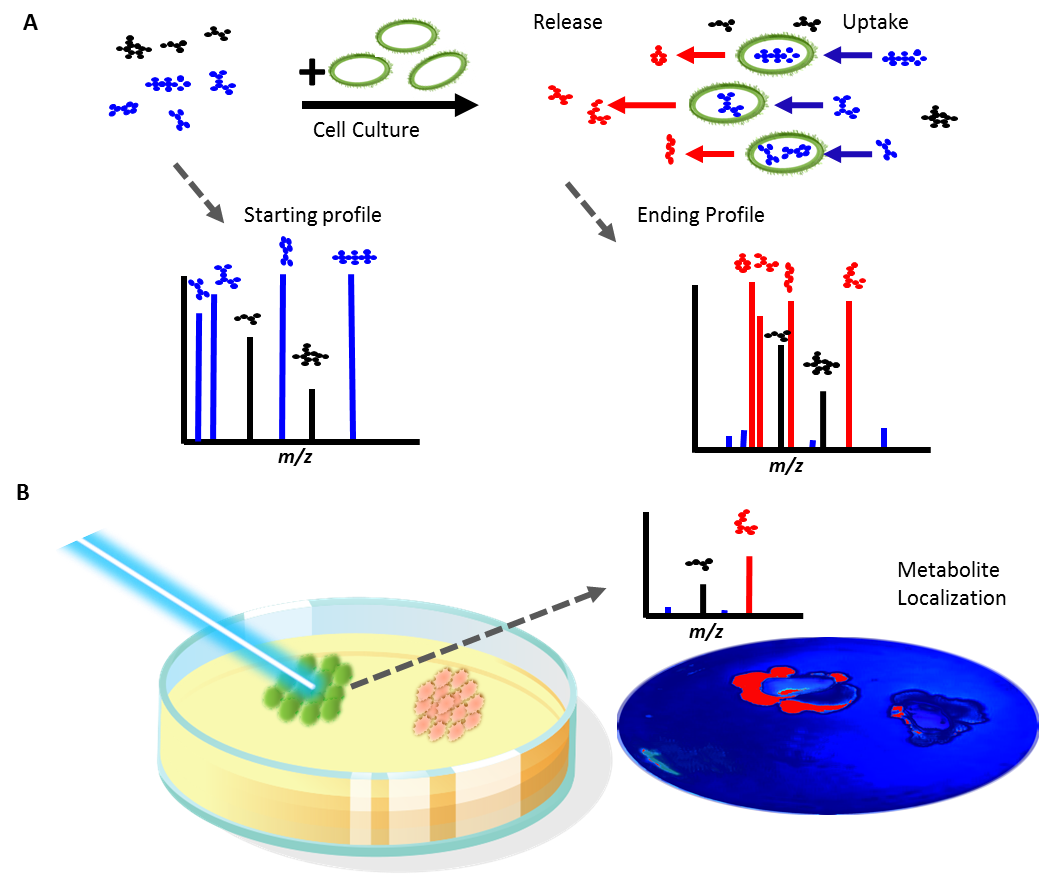
LCMS was used to determine the usage of compounds in soil-defined media (SDM) by different bacterial isolates. A significant amount of variation in compound depletion between isolates could be accounted for by rrn copy group and taxonomic group. Untargeted exometabolite profiling was also conducted to examine spent media from before and after growth experiments, revealing patterns of cross-feeding and metabolite production.
To learn more, check out the original article.
Reference:
Wang, Y., Wilhelm, R. C., Swenson, T. L., Silver, A., Andeer, P. F., Golini, A., Kosina, S. M., Bowen, B. P., Buckley, D. H., & Northen, T. R. (2022). Substrate Utilization and Competitive Interactions Among Soil Bacteria Vary With Life-History Strategies. Frontiers in Microbiology, 13. https://doi.org/10.3389/fmicb.2022.914472