Microfluidic metabolomics
Metabolomics and high throughput screening of molecular analytes are implemented in many areas of chemistry, medicine and industrial biotechnology. Increasing throughput is highly desirable for large-scale metabolomic studies and to streamline discovery of new chemical entities. Even with the recent development of high-throughput mass spectrometry, there are technical challenges for rapid sample preparation and processing. Advances in droplet microfluidics coupled with a massively parallel array-based approach can potentially be a solution.

To streamline sample preparation and processing in a high-throughput fashion for downstream surface-based MS analysis, we are developing a novel microfluidic system that can rapidly generate and manipulate hundreds of thousands of micro-droplets. The high density and small scale of arrayed droplets allow for parallel processing of sample directly on nanostructured MS surfaces and metabolomic screening to elucidate combinatorial complexity of interactions between molecules and environmental influences. This microfluidic system will potentially bridge the gap between analysis throughput and metabolite coverage of MS analysis.
We envision wide applications of this platform in screening of a massive library of enzymes as well as microbial community function to support bioenergy and sustainable bioproducts.
Nanostructure Initiator Mass Spectrometry (NIMS)
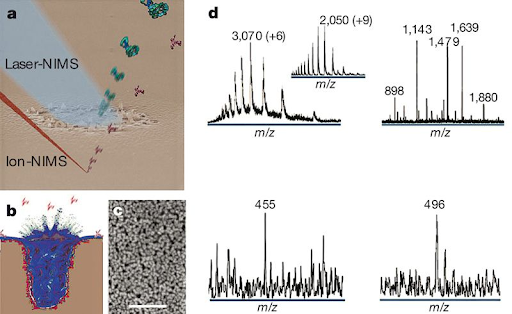
Nanostructure-Initiator Mass Spectrometry (NIMS) is a unique approach to mass spectrometry based on using liquid initiators, which are coated onto a nanostructured silicon surface [1].
An ionizing laser heats the surface, which results in the explosive vaporization of the trapped liquid coat as well as the adsorbed analyte molecules from surface applied samples. NIMS has been used to analyze a wide variety of analytes, including lipids, metabolites and for enzyme activity screenings [2,3].
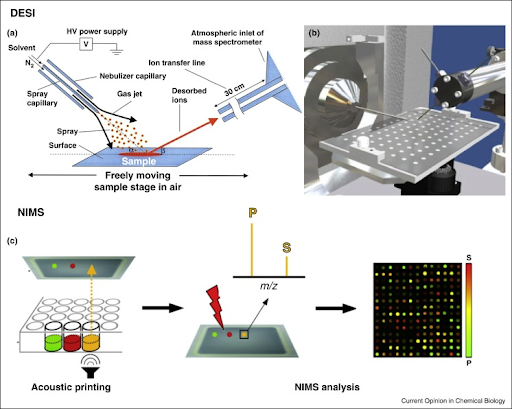
The sample throughput of NIMS can be significantly increased by increasing the number of samples spotted onto its surface. By using acoustic printing, we are able to spot 10,000 samples onto a NIMS surface and can reach a throughput of ~5 seconds per sample [4,5]. For example, this approach was used for functional characterization of glycosylhydrolases, where 10,000 different sample conditions were tested on enzymatic activity5.
NIMS can also be applied for metabolite analysis from tissue sections and microorganisms [6,7]. The replica-extraction-transfer (REX) technique was developed to allow for the transfer of metabolites directly from agar cultures onto NIMS surfaces and was used to investigate bacterial colony interactions [8].
Microbial community structure dynamics
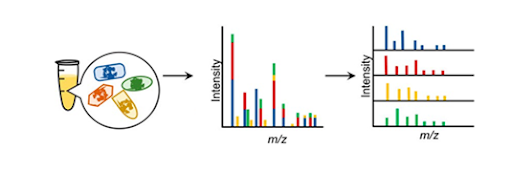
Although recent advances in molecular-based approaches have provided structural and functional relationships within microbial communities; variable copy numbers, dead vs live cell contribution, and abiotic pressures (ie; warming, wetting events, denitrification) can still attribute to read bias causing the appearance of changes in diversity measurements. Since bacterial communities actively respond to their environmental conditions, understanding the metabolite cross-talk between organisms is crucial in elucidating functional responses. To do this the composition of the community has to accurately be quantified to draw conclusions on who is accomplishing what.
To alleviate biases attributed to molecular techniques and deal with complex mixtures, we are utilizing lipid- and protein-based markers as fingerprint-like signatures of bacteria to construct a biomarker database for screening simple microbial communities. This allows for simultaneous elucidation of community structure and monitoring of nutrient cycling in response to external stimuli.
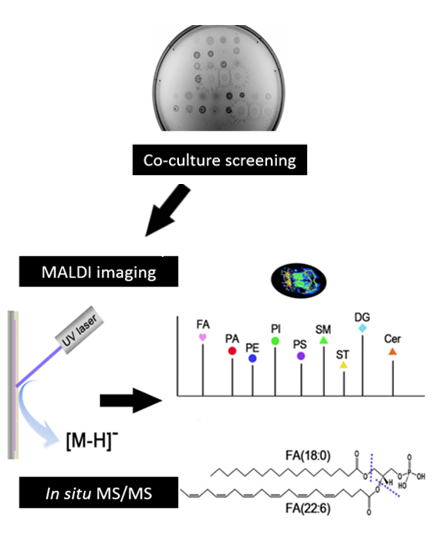
Profiling capabilities:
- Metabolomics/Metabolite Atlas
- REX-NIMS and Spray-Coat Imaging MS
- SIP of labeled nutrients
This approach can help provide a direct link between structure and function of a microbiome to dynamic environmental stressors. With the growing changes occurring as a result of climate change, land development, subsurface contamination, and etc, being able to link community response to direct environmental conditions will ultimately help in the promotion and sustainability of our vital ecosystems.
